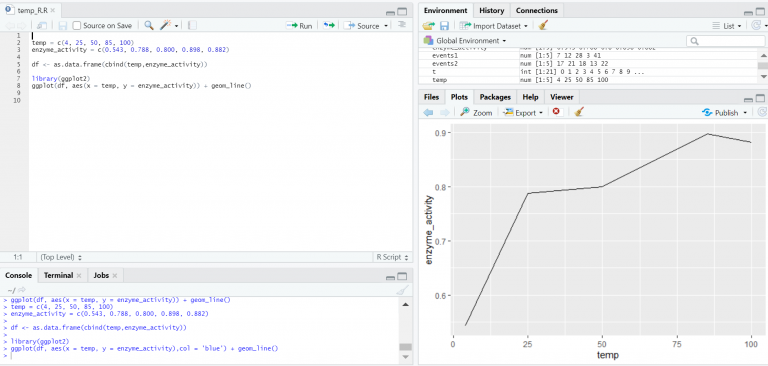

Nodes in the vicinity of a starting point can quickly be traversed giving the user the possibility to not only retrieve data but also perform fast analysis of these neighbour networks. No denormalisation is required so data can be stored in its natural form. Storing Reactome data in this form has many benefits.
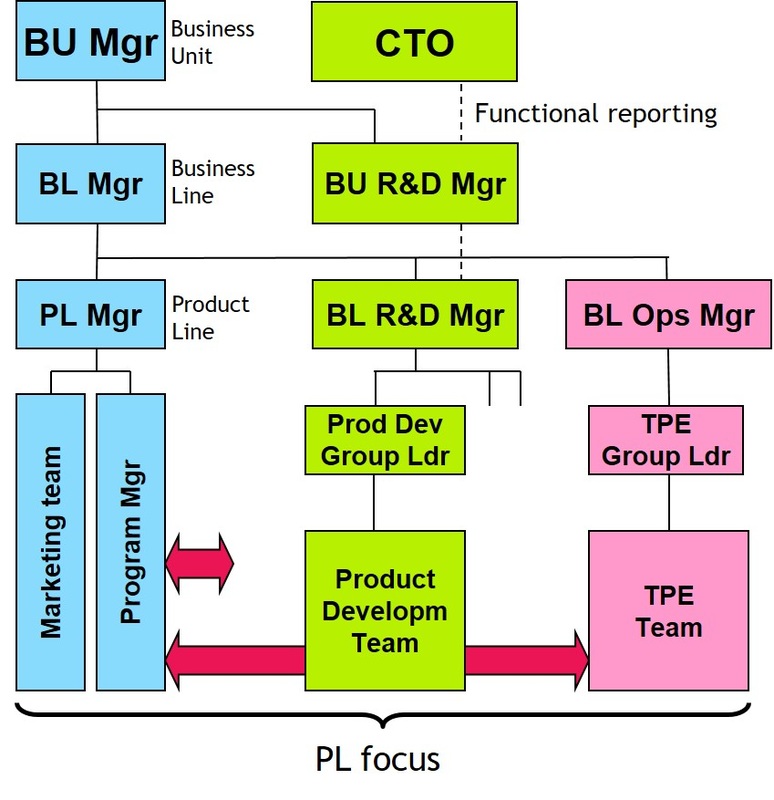
Graph database technology is an effective tool for modelling highly connected data. In order to overcome these problems the Reactome database is imported in Neo4j, creating one large interconnected graph. Due to the schema-based approach, relational databases are limited in how information is stored and thus are difficult to scale for new requirements. Queries across the pathway knowledgebase are composed by a number of expensive join operations resulting in poor performance and a hard-to-maintain project. Retrieving, and especially analysing such complex data becomes tedious when using relational databases. The Reactome Graph provides an intuitive way for data retrieval as well as interpretation and analysis of pathway knowledge. This amounts to millions of interconnected terms naturally forming a graph of biological knowledge. In Reactome, these processes are systematically described in molecular detail to generate an ordered network of molecular transformations (Fabregat et al. At the cellular level, life is a network of molecular reactions.
